Research
|
The molecular basis for generating cellular diversity
Embryonic patterning kicks off by initiating cellular divergence. The two cell fate decisions during preimplantation embryo development are each textbook examples of different heterogeneity-generating mechanisms, which are widely employed in nature. The first divergence arises from cell polarization followed by asymmetric cell division, yielding different daughter cell types. The second involves an initially uniform population of cells differentiating in a random “salt and pepper” pattern, followed by sorting out into distinct compartments. We are interested in the molecular mechanisms underlying the initiation of these events and seek to answer questions such as how polarization is initiated and inherited; and what the molecular determinants are for heterogeneous cellular response? |
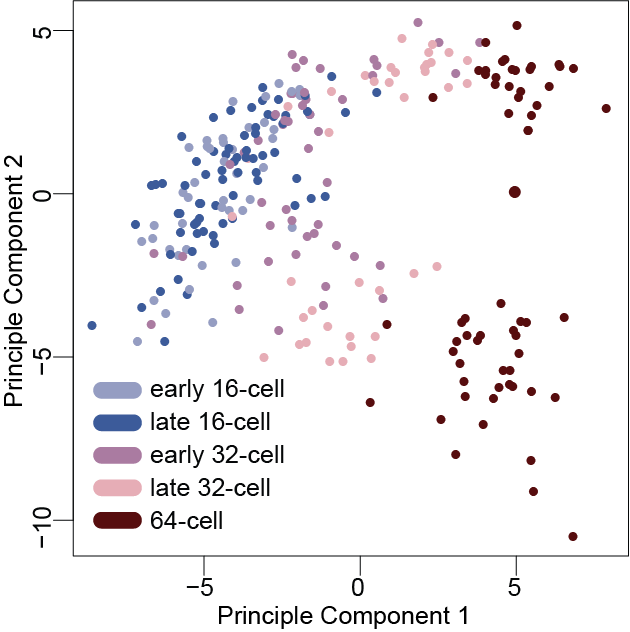
Transcriptional divergence emerging during trophectoderm and inner cell mass segregation revealed by single-cell RNA sequencing (Posfai et al 2017 eLife)
|
|
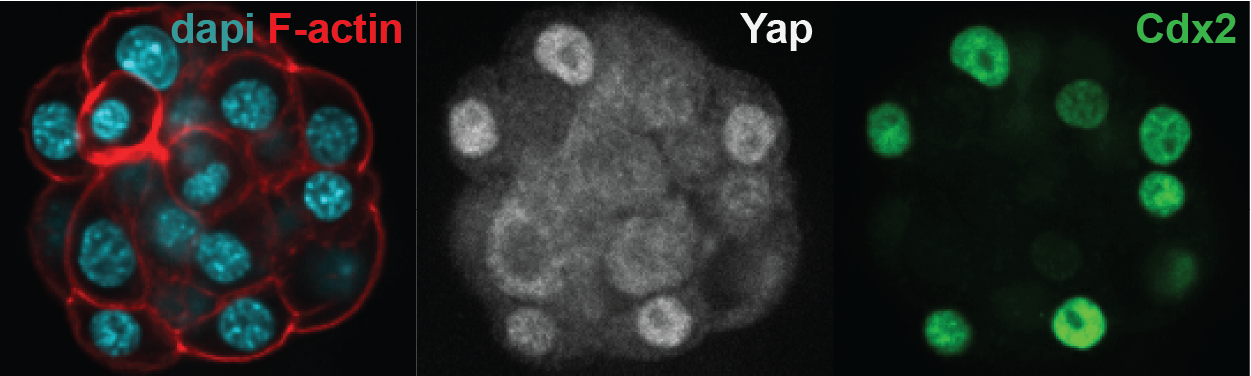
Differential Yap localization in the trophectoderm and the inner cell mass. Nuclear Yap co-localizes with the trophectoderm-specific transcription factor Cdx2.
|
The role of mechanical forces in shaping cell fate decisions in the embryo
Cell fate-specific signaling networks and gene expression are not only determined by the biochemical signals a cell receives, but are also controlled by the mechanical inputs a cell experiences. Mechanical forces are obviously at play during preimplantation embryo development – cells use the actomyosin cytoskeleton and adhesion molecules to generate and transmit forces, which result in embryo compaction, dynamic cell rearrangements and blastocyst cavity formation. We are specifically interested in how physical forces are relayed into cell fate decisions and how they complement known biochemical signaling activities. |
|
Mechanisms of totipotency in vivo and in vitro
Totipotency – the unbiased ability to generate both embryonic and extraembryonic cell types - has only unequivocally been shown to exist during the first few cell divisions of early mouse development. Surprisingly, very little is known about the molecular mechanisms controlling this unique state in mammals. Our goal is to identify the minimal gene regulatory network responsible for conferring such limitless potential by using a combination of bioinformatical approaches and harnessing our advances in CRISPR-based methods to probe gene function during early development. Additionally, guided by lessons from our embryo studies, we aim to establish a novel stem cell type with unbiased differentiation potential. These stem cells will serve as a source for generating synthetic embryos, which will in turn allow us to gain crucial developmental insights with the use of only in vitro substrates and will be a starting point for learning how to engineer a group of cells into more complex multicellular structures, such as tissues or organs. |
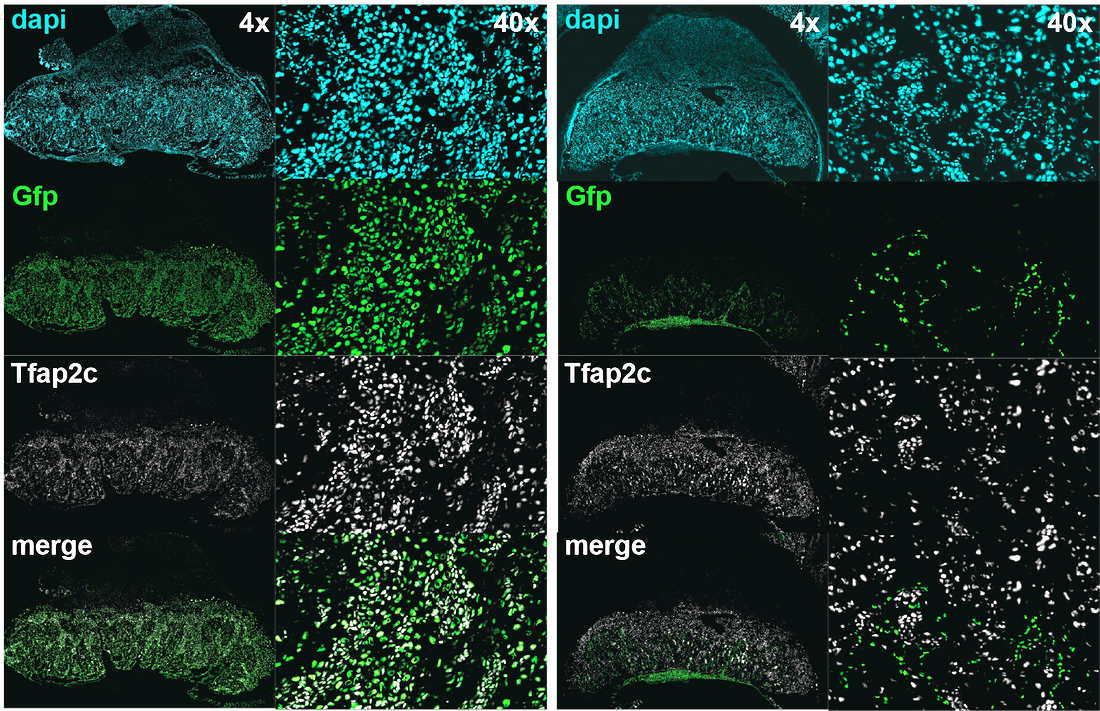
How do you test the developmental potential of a cell? By diligent lineage analysis in chimeras generated with a host embryo and the cell of interest. Images show different cell lineages present in the placenta, revealed by tetraploid complementation. Trophoblast (left panel) and embryonic (right panel) components of the placenta in GFP.
|